A package for efficient computations of standard clustering comparison measures
Installation
Stable version on the CRAN.
install.packages("aricode")The development version is available via:
devtools::install_github("jchiquet/aricode")Description
Computation of measures for clustering comparison (ARI, AMI, NID and even the distance) are usually based on the contingency table. Traditional implementations (e.g., function adjustedRandIndex of package mclust) are in where
- is the size of the vectors the classifications of which are to be compared,
- and are the respective number of classes in each vectors.
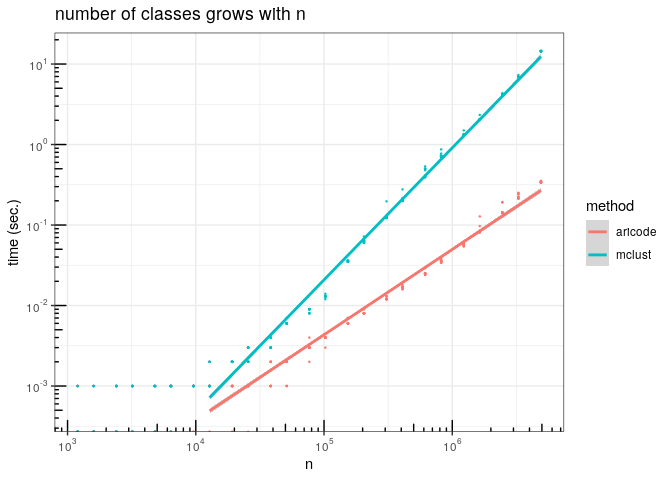
In aricode we propose an implementation, based on radix sort, that is in in time and space.
Importantly, the complexity does not depend on and . Our implementation of the ARI for instance is one or two orders of magnitude faster than some standard implementation in R.
Available measures and functions
The functions included in aricode are:
-
(A)RI: computes the (adjusted) Rand index -
MARI: computes the modified adjusted rand index as defined in Sundqvist et al, 2023 -
NID: computes the normalized information distance -
NMI: computes the normalized mutual information -
NVI: computes the the normalized variation information -
AMI: computes the adjusted mutual information -
Chi2: computes the Chi-square statistics -
Frobeniuscompute the Frobenius norm between two classification as defined in Arlot et al, 2019 -
entropy: computes the conditional and joint entropies -
compare_clustering: computes all clustering comparison measures at once -
sort_pairs: radix sort for pairs of elements